Section: New Results
Knowledge-based generalization of metabolic models
Participants : David James Sherman [correspondant] , Pascal Durrens, Razanne Issa, Anna Zhukova.
Large metabolic networks are hard to understand and curate, because the large number of detailed reactions, which are needed for accurate modeling and simulation, obscure the high-level structure of the reaction network. We defined knowledge-based methods that factor similar reactions into “generic” reactions in order to visualize a whole pathway or compartment, while maintaining the underlying model so that the user can later “drill down” to the specific reactions if need be[15] , [16] An implementation of this method is available as a Python library (see paragraph 5.3 ).
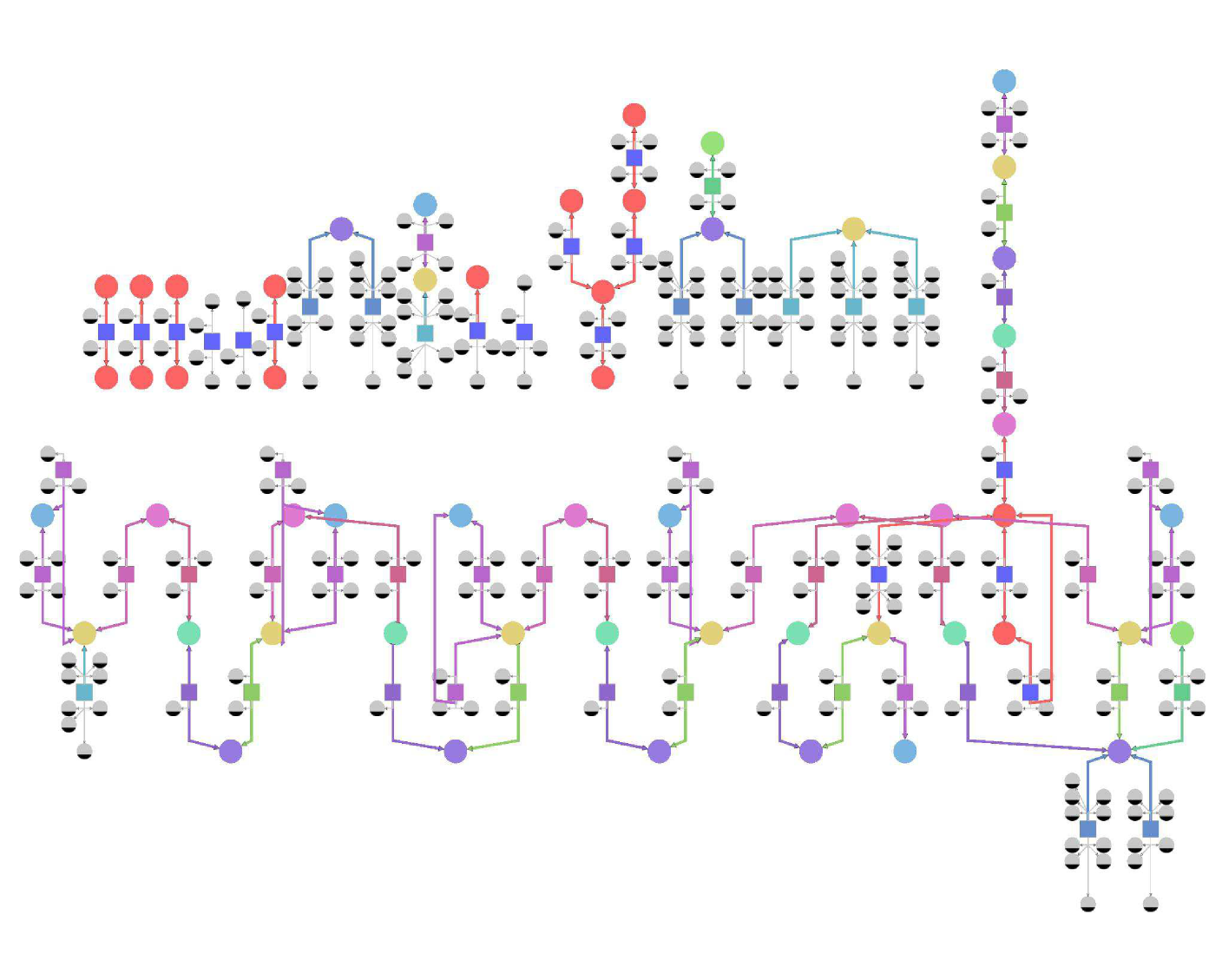
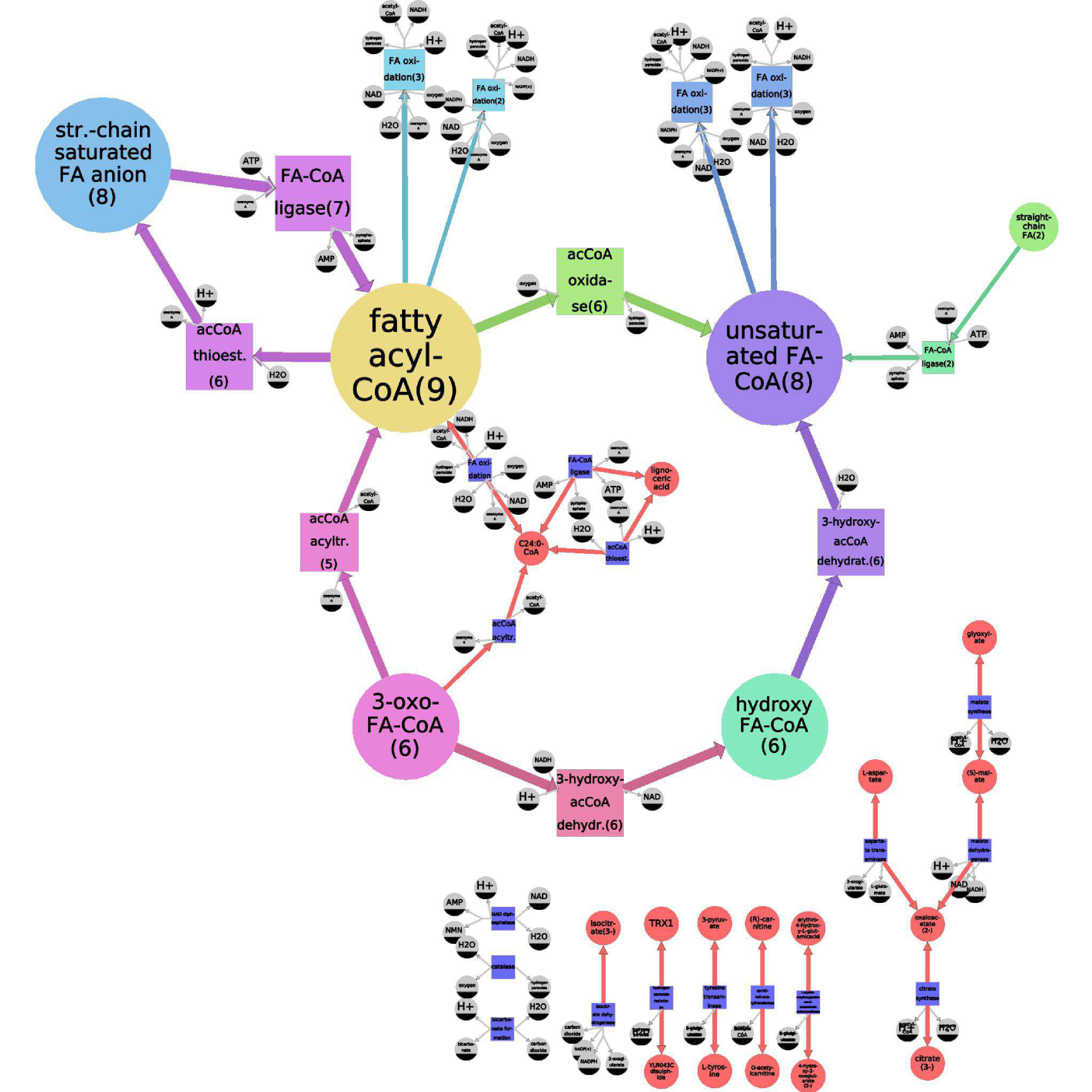
Figures 2 and 3 illustrate model generation for Yarrowia lypolitica fatty acid oxidation in the peroxisome. Molecular species are represented using SBGN notation: as circular nodes, and the reactions as square ones, connected by edges to their reactants and products. Ubiquitous species are of smaller size and colored gray. Non-ubiquitous species are divided into six equivalence classes, and coloured accordingly. The size of the model does not allow for readability of the species labels, thus we do not show them (figure 2 ).
The specific model is appropriate for simulation, because it contains all of the precise reactions. The generalized model is suited for a human, because it reveals the main properties of the model and masks distracting details. For example, the generalized model highlights the fact that there is a particularity concerning C24:0-CoA (stearoyl-CoA) (yellow): there exists a "shortcut" reaction (orange), producing it directly from another fatty acyl-CoA (yellow), avoiding the usual four-reaction beta-oxidation chain, used for other fatty acyls-CoA. This shortcut is not obvious in the specific model, because it is hidden among a plethora of similar-looking reactions.
We formally defined the generalization method in [15] and showed how to calculate it using a good approximation to an NP-complete set cover problem. The method was further validated in a collection of 1283 inferred models and revealed, on the one hand, a number of probable errors in the inferred models, and on the other hand, that there exist different families of generalization with a plausible link to different adaptive responses.
|
|